What are protein interactions?
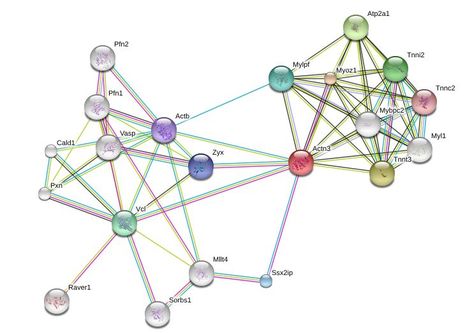
Interactions between proteins are important for the majority of biological functions. These interactions are often the result of proteins binding to each other and can be hard to characterize due to their complexity and dynamic nature. Scientists use a variety of methods such as yeast-two hybrids, TAP-tagging, and coimmunoprecipitation to identify these interactions and create networks as shown in Figure 1.
Yeast-two Hybrids
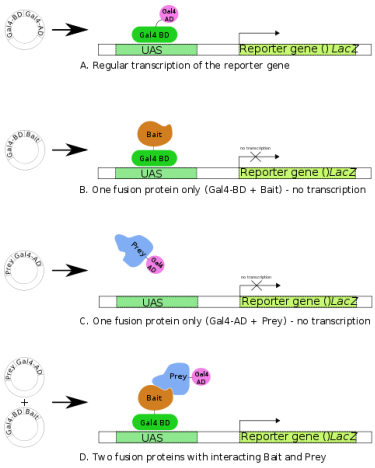
Yeast-two Hybrid (YTH) screens are used to detect interactions between two or more proteins and proteins and DNA. YTH screens work by fusing a bait protein to a DNA binding domain, and fusing a prey protein to an activation domain of a transcription factor (1). This will then turn on a reporter gene of interest (see Figure 2). Experiments using YTH screens demonstrate how transcription factors are capable of activating transcription without directly binding to one another.
TAP-tagging
Tandem Affinity Purification tags work by tagging the protein of interest so it binds with an IgG bead. The TAP tag is then cleaved at a specific site to allow the complex to bind to calmodulin coated beads (3). The resulting complex is free of any contaminants. TAP tag technology is very reliable and the results are very reproducible. This is in part because any bait protein attached to a specific complex will pull down the same protein as any other protein in that complex. (4).
Co-immunoprecipitation
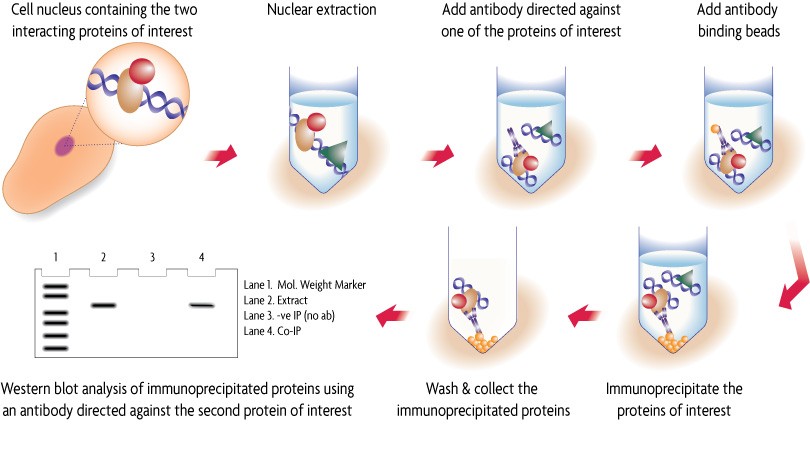
Co-immunoprecipitation (Co-IP) works by selecting an antibody that targets a known protein that is believed to be a member of a larger complex of proteins. The antibody-protein complex allows researchers to pull the entire protein complex out of solution and identify the other proteins bound in the complex (see Figure 3) (5). This works when the proteins involved in the complex bind to each other tightly, making it possible to pull multiple members of the complex out of solution by latching onto only one.
Analysis
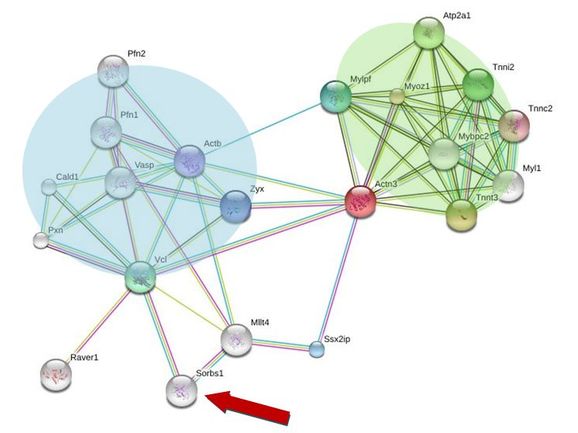
In order to analyze protein interactions I used STRING to create the network shown below. STRING gives you the ability to zoom in on proteins that more directly interact with the protein of interest or to zoom out a level to see which proteins are a step out from the protein of interest. I searched for ACTN3 (shown as the red circle in Figure 4) and found a number of different proteins that interact with it.
Discussion
When STRING returns with your protein network it also provides a brief synopsis on the function of the related proteins. I looked at these protein functions to determine if there was anything unusual about the interactions. The proteins shaded by the blue box (Figure 4) are needed for cytoskeletal rearrangements and the actin/actinin binding. The proteins shaded by the green box are needed for muscle contraction in fast twitch muscles and myosin binding. Both of these are rather expected since researchers understand the role ACTN3 plays in bind actin and myosin filaments together. However, the interaction of ACTN3 with SORBS1 (red arrow in Figure 4) was interesting to me because SORBS1 plays a role in tyrosine phosphorylation and has been linked to insulin dependent glucose transport out of cells. I thought that this may be a potential link to ACTN3's possible role in metabolism.
References
1 Suter, B., Kittanakom, S., Stagljar, I. (2008) Two-hybrid technologies in proteomics research. Current Opinion in Biotechnology. 2008 19(316).
2. Wikipedia.(2007) Yeast-Two Hybrid Screening. http://en.wikipedia.org/wiki/File:Two_hybrid_assay.svg
3. Collins, M. O., Choudhary, J. S. (2008) Mapping multiprotein complexes by affinity purification and mass spectrometry. Current Opinion in Biotechnology. 2008 19(324).
4. Gavin, AC., Bosche, M., Krause, R., Grandi, P., Marzioch, M., Bauer, A., Schultz, J., Rick, J. M., Michon A-M., Cruciat, C-M., Remor, M., Hofert, C., Schelder, M., Brajenovic, M., Ruffner, H., Merino, A., Klein, K., Hudak, M., Dickson, D., Rudi, T., Gnau, V., Gauch, A., Bastuck, S., Huhse, B., Leutwein, C., Heurtier, M-A., Copley, R., Edelmann, A., Querfurth, E., Rybin, V., Drewes, G., Raida, M., Bouwmeester, T., Bork, P., Seraphin, B., Kuster, B., Neubauer, G., Superti-Furga, G. (2002)
Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature. 415(141).
5. Sino Biological Inc. 2007. Co-immunoprecipitation Background and Protocol. http://www.assay-protocol.com/Immunology/Co-IP
6. Active Motif. Nuclear Complex Pro-IP kit. Description. http://www.activemotif.com/catalog/25/nuclear-complex-co-ip-kit
2. Wikipedia.(2007) Yeast-Two Hybrid Screening. http://en.wikipedia.org/wiki/File:Two_hybrid_assay.svg
3. Collins, M. O., Choudhary, J. S. (2008) Mapping multiprotein complexes by affinity purification and mass spectrometry. Current Opinion in Biotechnology. 2008 19(324).
4. Gavin, AC., Bosche, M., Krause, R., Grandi, P., Marzioch, M., Bauer, A., Schultz, J., Rick, J. M., Michon A-M., Cruciat, C-M., Remor, M., Hofert, C., Schelder, M., Brajenovic, M., Ruffner, H., Merino, A., Klein, K., Hudak, M., Dickson, D., Rudi, T., Gnau, V., Gauch, A., Bastuck, S., Huhse, B., Leutwein, C., Heurtier, M-A., Copley, R., Edelmann, A., Querfurth, E., Rybin, V., Drewes, G., Raida, M., Bouwmeester, T., Bork, P., Seraphin, B., Kuster, B., Neubauer, G., Superti-Furga, G. (2002)
Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature. 415(141).
5. Sino Biological Inc. 2007. Co-immunoprecipitation Background and Protocol. http://www.assay-protocol.com/Immunology/Co-IP
6. Active Motif. Nuclear Complex Pro-IP kit. Description. http://www.activemotif.com/catalog/25/nuclear-complex-co-ip-kit