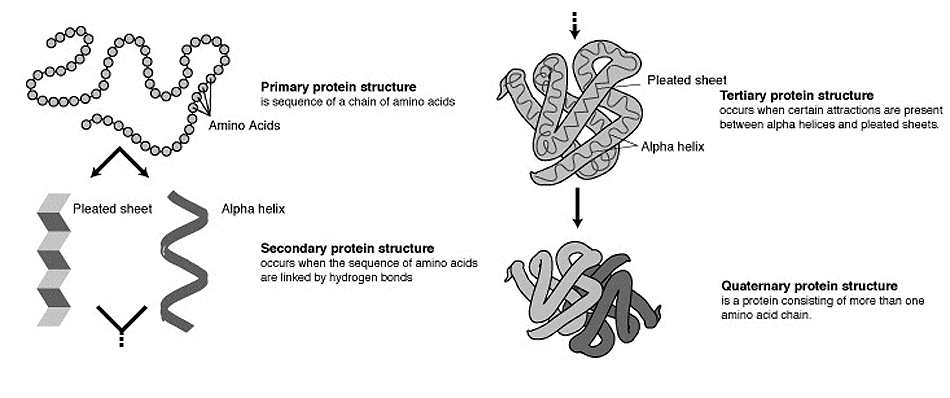
What are protein domains and motifs?
Protein motif refers to the “building blocks” of the protein,
the actual amino acid sequence of the polypeptide chain. This sequence is also
known as the primary structure of the protein. When several sequences are
combined together they can create a functional unit known as a domain (Figure 1). Protein
domains are components of a greater protein that is capable of functioning
singularly without the entirety of the protein structure. The protein domain
may be a result of the physical arrangement of the protein in space and confer
specific functions of the protein. Some of these functions include binding
sites, catalytic capabilities, and signal transduction.
These
domains are often similar or conserved across many different species and using
the domains proteins can be grouped into different families. Because the
domains have similar functions despite the species it can give us information
about the origin of protein in terms of evolution. Most proteins are
multi-domain proteins, which means they have multiple domains within their
structure. Using SMART it is possible to see that ACTN3 is one of these multi-domain proteins (Figure 3). The multiple domains
probably existed in independent structures at some point but have evolved
together onto the same protein to perform a specific protein function. The
multiple domains came together from the transposition of mobile genetic
elements, large scale gene mutations -
deletions, duplication, translocations etc. - homologous
recombination and DNA polymerase replication errors.
Analysis
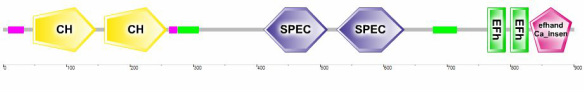
Figure 3. SMART analysis shows the major domains in the human ACTN3 proteins. The notable functional domains are the CH domain, Spec domain, Efh domain, and elfhand Ca_insen domain.
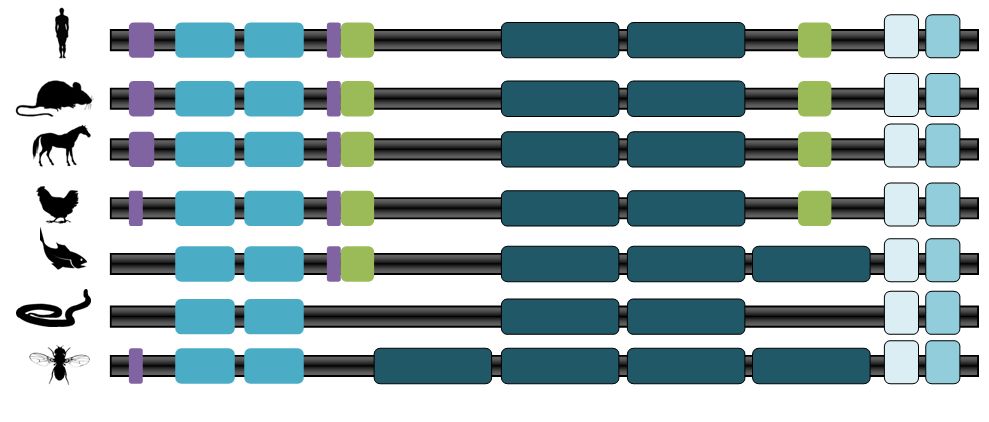
The alpha-actinin family of proteins are highly conserved across several different species, including non-vertebrates. This is evident by looking at the protein homology (see homology information) and by the fact that ACTN3 has been conserved since it's divergence from ACTN2 approximately 300 million years ago (1). Therefore it is logical that the important domains would be conserved across most species (Figure 2). Even the length of protein is well conserved, differing only by about 4 amino acids between the human protein (897 amino acids) and the nematode protein (893 amino acids).
One important and well conserved domain is the Calponin Homology (CH) Domain which is the actin binding domain (2). Since ACTN3 is a specialized skeletal muscle actin binding protein the CH Domain is essential for correct ACTN3 functioning. The EF hand and the Ca2+ binding domain are also highly conserved amongst species, possibly because Ca2+ binding is necessary for muscle contraction and relaxation and ACTN3 works with actin to contract muscles.
The domains that are most variable in the ACTN3 protein are the spectrin reapeats, the low complexity region, and the coiled coil region. Both the spectrin repeats and the coiled coil regions are related to the protein structure. The mammal proteins have similar protein structure domain and therefore a well conserved protein structure past the divergence of the mammal line. It is interesting to me that the zebrafish has a 3rd spectrin repeat in place of a coiled coil region and I wonder how this changes the ACTN3 structure and interaction with actin.
http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1180686/
One important and well conserved domain is the Calponin Homology (CH) Domain which is the actin binding domain (2). Since ACTN3 is a specialized skeletal muscle actin binding protein the CH Domain is essential for correct ACTN3 functioning. The EF hand and the Ca2+ binding domain are also highly conserved amongst species, possibly because Ca2+ binding is necessary for muscle contraction and relaxation and ACTN3 works with actin to contract muscles.
The domains that are most variable in the ACTN3 protein are the spectrin reapeats, the low complexity region, and the coiled coil region. Both the spectrin repeats and the coiled coil regions are related to the protein structure. The mammal proteins have similar protein structure domain and therefore a well conserved protein structure past the divergence of the mammal line. It is interesting to me that the zebrafish has a 3rd spectrin repeat in place of a coiled coil region and I wonder how this changes the ACTN3 structure and interaction with actin.
http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1180686/