Chemical Genetics
Chemical genetics uses small molecules to directly change the way proteins work by interacting with them directly. This is compared to indirectly changing proteins through manipulating the protein building blocks, genes. The term chemical genetics refers to the chemistry used to generate the small molecules to interact with the proteins (1). Chemical genetics can be used to identify which proteins are involved in different biological processes and to understand how these proteins perform their functions at a molecular level (1). The small molecules can have a variety of effects on a protein, including rendering them non-functional or aiding in proper folding (and therefore increasing functionality of an inherently mis-folded, non-functional protein).
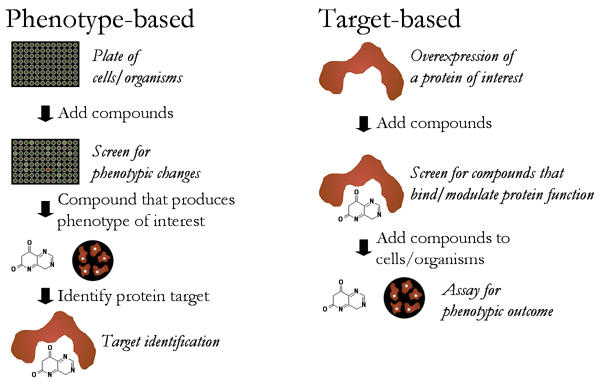
There are two main approaches when it comes to chemical genetics (Figure 1). One approach, the forward chemical genetic screen, starts with a phenotype or characteristic and works to identify the protein or proteins responsible. The second approach, reverse chemical genetic screen, scientists start with a specific protein and try to determine its function by studying the change in phenotype when the protein interacts with the small molecules (2).
There are two main approaches when it comes to chemical genetics (Figure 1). One approach, the forward chemical genetic screen, starts with a phenotype or characteristic and works to identify the protein or proteins responsible. The second approach, reverse chemical genetic screen, scientists start with a specific protein and try to determine its function by studying the change in phenotype when the protein interacts with the small molecules (2).
Small molecules offer the potential to treat specific diseases by targeting the over-expressed or incorrectly functioning proteins. ChemBank is a chemical library database that organizes small molecules and the proteins they interact with. Unfortunately, there were no identified small molecules that have been shown to interact with ACTN3.
RNA interference
RNA interference (RNAi) is so named because it interferes with the function of the RNA to change the organisms phenotype. RNAi uses double stranded RNA to silence expression of a gene by targeting messenger RNA. When mRNA is destroyed, the gene product cannot be translated into a functional protein, effectively silencing the expression of a gene (3). RNAi is a powerful tool for targeting a specific gene and analyzing the resulting phenotype because it gives an otherwise healthy animal a diseased phenotype. Using various RNAi databases, a researcher can study the results of past experiments in order to decide which organism to use for current research. Most experiments are done in model organisms such as fruit flies, zebrafish, and C. elegans.
I searched for ACTN3 in different databases such as RNAiDB and Wormbase but I was unable to find any past experiments documenting RNAi use when studying ACTN3. This is probably because research surrounding ACTN3 focuses more on the phenotypes expressed in humans and the mutation that results in the different phenotypes are not found in animals. Some researchers have created a mouse model which mimics the effect of the null ACTN3 phenotype, this suggests that mice are currently the best model organism to study ACTN3.
I searched for ACTN3 in different databases such as RNAiDB and Wormbase but I was unable to find any past experiments documenting RNAi use when studying ACTN3. This is probably because research surrounding ACTN3 focuses more on the phenotypes expressed in humans and the mutation that results in the different phenotypes are not found in animals. Some researchers have created a mouse model which mimics the effect of the null ACTN3 phenotype, this suggests that mice are currently the best model organism to study ACTN3.
DNA Microarrays
When cells are translating genetic material into proteins they form an intermediary called messenger RNA (mRNA) molecules, these mRNA can be used to determine how many genes are actively creating proteins, or being expressed, to perform essential cell functions in response to changing needs (4). Researchers use DNA Microarrays to study the expression levels of hundreds or thousands of genes within a cell. The Mircoarrays use small supports unto which the sequences of different genes are immobilized at specific sites. mRNA collected from cells then binds to its original DNA sequence on the Microarray plate (4). Using a computer, researchers are able to calculate the amount of mRNA bound to each spot and create a profile of gene expression in the cell (Figure 2). There are large databases of microarrays, such as GEO, that have been created and contain information about gene expression in various animals, tissues, and disease states. Not entirely surpringly, there is not very much information about ACTN3 in GEO. Probably due to similar reasons as stated above, there simply isn't much data on the actual gene function of ACTN3.
References
1. Howard Hughes Medical Institute. What is chemical genetics? Genomics and Chemical Genetics. Retreived from http://www.hhmi.org/biointeractive/genomics/poster_a2.html
2. Plant Systems Biology. Chemical Genetics in General. Retrieved from http://www.psb.ugent.be/roots-projects/157-chemical-genetics
3. Nature. RNA interference. Retrieved from http://www.nature.com/nrg/multimedia/rnai/index.html
4. National Center for Biotechnology Information. (2007). Microarrays: Chipping away at the mysteries of science and medicine. Retrieved from http://www.ncbi.nlm.nih.gov/About/primer/microarrays.html
2. Plant Systems Biology. Chemical Genetics in General. Retrieved from http://www.psb.ugent.be/roots-projects/157-chemical-genetics
3. Nature. RNA interference. Retrieved from http://www.nature.com/nrg/multimedia/rnai/index.html
4. National Center for Biotechnology Information. (2007). Microarrays: Chipping away at the mysteries of science and medicine. Retrieved from http://www.ncbi.nlm.nih.gov/About/primer/microarrays.html