What is protein homology?
Homology in biology is a concept of quality, defining the relationship between two things. Protein homology studies proteins in different species to determine how related they are. It is strictly defined as two proteins having a common ancestral protein (1). Over time, different species evolve but the protein stays the same within the new species. If the protein retains the same function during this process they can also be considered orthologous proteins (2). The protein is then passed from parent to offspring in what is known as vertical homology (3). A second way homologous proteins can be found in two species is a bit more complicated. If the gene coding for the protein is duplicated in an organism then it will be present twice in the genome. Then a speciation event occurs, and one copy of the gene is preserved in each new species (4). The two proteins encoded in the genomes can have similar functions, but selective pressures could also result in different functionality in one copy.
Why do we study homology?
In
human genetic research, it is important to identify homologous proteins in
model organisms in order to conduct research to discover the true nature of the
protein. Protein homology can give us information about which model organisms can be used to study the protein or gene in question. There are a number of ethical concerns about performing studies on humans, as well as fiscal and temporal issues for scientists. These models make it possible to study proteins more thoroughly than would be possible in otherwise. Additionally, if a protein is conserved in form and function, it suggests that the protein is involved in a pathway that is important or necessary to sustain a certain level of functioning (4).
What do you think?
Now that you know more about homology it's time to make an educated guess about the ACTN3 protein.Finding the Homologs of ACTN3
Homologs were found using NCBI's Basic Local Alignment Search Tool (BLAST) program. BLAST searches for regions of similarity between sequences across multiple genomes. Using algorithms and statistics BLAST creates alignments and predict the percent of identical sequences. This information can be used to find relationships between sequences. To use BLAST to find homologs, the sequence of interest is entered into the program and a list of potentially homologous genes in other species (bases on how identical the sequences are) is generated.
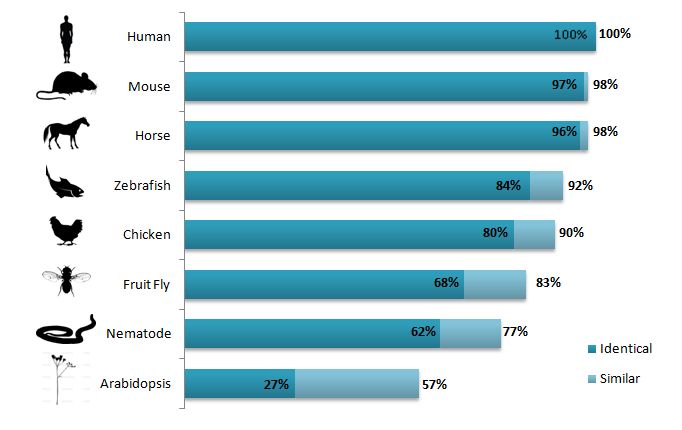
BLAST is also great for a detailed analysis of the homologous sequences. It can provide information about how significant the homology is and how closely related the sequences are. The E value generated by BLAST (listed below) describes how expected it is that the sequences match may have occurred by chance. The lower the E Value, the more significant the homology is. In addition to listing how identical the sequences are, BLAST can also give information about how similar the sequences are. Proteins are created from different chemical codes called amino acids. When the amino acid sequence for the reference protein (in this case Human ACTN3) matches exactly with another sequence it is identical. When the amino acids don't match exactly but the amino acid in the reference proteins have similar chemical properties (and therefor a similar function) to an amino acid in the other sequence they are referred to as similar. Because similarity has a more flexible definition sequences are often more similar than they are identical (see Figure 2 for comparisons).
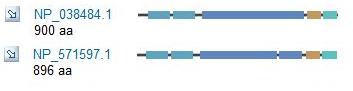
Another program, Homologene, can be used to initially generate a list of homologous proteins and give a visual depiction of any differences in conserved domains. Unfortunately, Homologene did not have the human ACTN3 in it's database. It did, however, have a number of other species' sequences that were homologous to the ACTN3 sequence.
BLAST is also great for a detailed analysis of the homologous sequences. It can provide information about how significant the homology is and how closely related the sequences are. The E value generated by BLAST (listed below) describes how expected it is that the sequences match may have occurred by chance. The lower the E Value, the more significant the homology is. In addition to listing how identical the sequences are, BLAST can also give information about how similar the sequences are. Proteins are created from different chemical codes called amino acids. When the amino acid sequence for the reference protein (in this case Human ACTN3) matches exactly with another sequence it is identical. When the amino acids don't match exactly but the amino acid in the reference proteins have similar chemical properties (and therefor a similar function) to an amino acid in the other sequence they are referred to as similar. Because similarity has a more flexible definition sequences are often more similar than they are identical (see Figure 2 for comparisons).
Another program, Homologene, can be used to initially generate a list of homologous proteins and give a visual depiction of any differences in conserved domains. Unfortunately, Homologene did not have the human ACTN3 in it's database. It did, however, have a number of other species' sequences that were homologous to the ACTN3 sequence.
Discussion
The ACTN3 protein is very similar in most of the model organisms, which suggests that it is a well conserved protein throughout evolution (Figure 2). The mammals are almost completely identical and the other vertebrae are 80% identical and still 90% similar. The invertebrates begin to differ the most but even the nematode ACTN3 protein is 77% similar which is pretty remarkable when you think about how different humans appear from nematodes and fruit flies!
Homologous Protein Reference Numbers
|
Humans(Homo sapiens)-alpha actinin 3
Accession Number: NP_001245300.1 GI Number: 385648244 FASTA Horse (Equus caballus)-alpha actinin 3 Accession Number:NP_001157341.1 GI Number: 255522877 FASTA E value: 0.0 Max identical: 96% % Similar: 98% Chicken (Gallus gallus)-alpha actin 2 Accession Number:NP_990654.1 GI Number: 46048687 FASTA E value: 0.0 Max identical: 80% % Similar: 90% Nematode (Caenorhabditis elegans) -Protein ATN1 Accession Number:NP_506128.1 GI Number: 17565034 FASTA E value: 0.0 Max identical: 62% % Similar: 77% |
Mouse (Mus musculus)-alpha actinin 3
Accession Number:NP_038484.1 GI Number: 7304855 FASTA E value: 0.0 Max identical: 97% % Similar: 98% Zebrafish (Danio rerio)- alpha actinin Accession Number: AAN77132.1 GI Number: 25992501 FASTA E value: 0.0 Max identical: 84% % Similar: 92% Fruit Fly (Drosophila melanogaster)-alpha actinin Accession Number:XP_002057359.1 GI Number: 195397485 FASTA E value: 0.0 Max identical: 68% % Similar: 83% Arabidopsis - Calcium binding protein CML20 Accession Number: NP_190605.1 GI Number: 15229732 FASTA E value: 9e-12 Max identical: 27% % Similar: 57 % |
References
(1) Reeck, G. R., de Haen, C, Teller, D. C., Doolittle, R. F., Fitch, W. M., Dickerson, R. E., Chambon, P., McLachlan, A. D., Margoliash, E., Jukes, T. H., Zuckerandl E. (1987)
“Homology” in proteins and nucleic acids: a terminology muddle and a way out of it. Cell, 1987(50), 667. doi: 10.1016/0092-8674(87)90322-9
(2) Delsuc F, Brinkmann H, Philippe H. Phylogenomics and the reconstruction of the tree of life. Nat Rev Genet. 2005 May;6(5):361-75. Review. PubMed PMID: 15861208.
(3) Fitch W. (1970) Distinguishing homologous from analogous proteins. Systematic Zoology, 1970(19), 99. Retrieved from: http://www.jstor.org/stable/2412448
(4) Studer, R. A., Robinson-Rechavi, M. (2005)
How confident can we be that orthologs are similar, but paralogs differ? Trends in Genetics, 2005(25), 210. doi: 10.1016/j.tig.2009.03.004
(3) Brody, Thomas B., PhD. "Evolutionarily Conserved Developmental Pathways." The Interactive Fly. Society for Developmental Biology, 10 Feb. 2012. Web. 15 Feb. 2013. <http://www.sdbonline.org/fly/aimain/aadevinx.htm>.
“Homology” in proteins and nucleic acids: a terminology muddle and a way out of it. Cell, 1987(50), 667. doi: 10.1016/0092-8674(87)90322-9
(2) Delsuc F, Brinkmann H, Philippe H. Phylogenomics and the reconstruction of the tree of life. Nat Rev Genet. 2005 May;6(5):361-75. Review. PubMed PMID: 15861208.
(3) Fitch W. (1970) Distinguishing homologous from analogous proteins. Systematic Zoology, 1970(19), 99. Retrieved from: http://www.jstor.org/stable/2412448
(4) Studer, R. A., Robinson-Rechavi, M. (2005)
How confident can we be that orthologs are similar, but paralogs differ? Trends in Genetics, 2005(25), 210. doi: 10.1016/j.tig.2009.03.004
(3) Brody, Thomas B., PhD. "Evolutionarily Conserved Developmental Pathways." The Interactive Fly. Society for Developmental Biology, 10 Feb. 2012. Web. 15 Feb. 2013. <http://www.sdbonline.org/fly/aimain/aadevinx.htm>.