What is DNA homology?
Homology in biology is a concept of quality, defining the relationship between two things. Protein homology studies proteins in different species to determine how related they are. It is strictly defined as two proteins having a common ancestral protein (1). Over time, different species evolve but the gene stays the same within the new species. If the gene retains the same function during this process they can also be considered orthologous (2). The gene is then passed from parent to offspring in what is known as vertical homology (3). A second way homologous proteins can be found in two species is a bit more complicated. If the gene is duplicated in an organism then it will be present twice in the genome. Then a speciation event occurs, and one copy of the gene is preserved in each new species (4).
Discussion
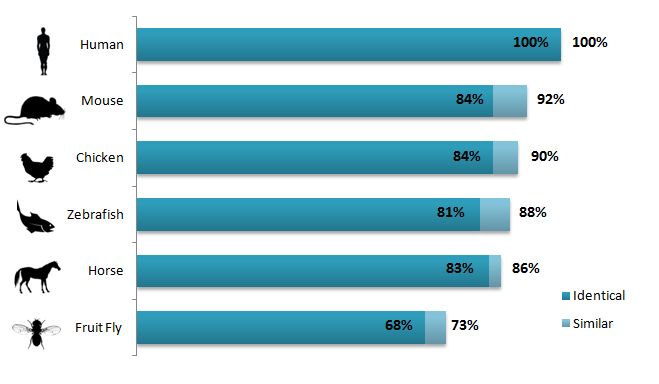
The ACTN3 gene is very similar in most of the model organisms, which suggests that it is a well conserved protein throughout evolution (Figure 1). The percent similar and the percent identical are very close in all of the organisms, which suggests that the parts that match the human ACTN3 gene are a majority of what is similar in the other organism's genes. I thought that it was interesting that the gene homology differs than the protein homology (as seen here). The horse gene is not very similar to the human gene but the horse protein is more similar. This is probably a result of events during transcription and translation, or the protein formation, that saves the parts of the gene that are similar to humans and gets rid of the parts of the gene that are different than humans. I was unable to have as thorough of a comparison with invertebrate model organism with the gene because I couldn't find the ACTN3 DNA sequence of a number of them. However, the fruit fly does seem to fit the general theme that the gene is more similar to a human than the protein.
|
Humans(Homo sapiens)-alpha actinin 3
Accession Number: NC_000011.9 GI Number: 385648244 FASTA Horse (Equus caballus)-alpha actinin 3 Accession Number: NC_009155.2 GI Number: 255522877 FASTA E value: 0.0 Max identical: 83% % Similar: 86% Chicken (Gallus gallus)-alpha actin 4 Accession Number: NM_205126.1 GI Number: 46048687 FASTA E value: 0.0 Max identical: 84% % Similar: 90% |
Mouse (Mus musculus)-alpha actinin 3
Accession Number: NC_000085.6 GI Number: 7304855 FASTA E value: 0.0 Max identical: 84% % Similar: 92% Zebrafish (Danio rerio)- alpha actinin Accession Number: NC_007132.5 GI Number: 25992501 FASTA E value: 0.0 Max identical: 81% % Similar: 88% Fruit Fly (Drosophila melanogaster)-alpha actinin Accession Number: NT_033777.2 GI Number: 195397485 FASTA E value: 0.0 Max identical: 68% % Similar: 73% |
References
(1) Reeck, G. R., de Haen, C, Teller, D. C., Doolittle, R. F., Fitch, W. M., Dickerson, R. E., Chambon, P., McLachlan, A. D., Margoliash, E., Jukes, T. H., Zuckerandl E. (1987)
“Homology” in proteins and nucleic acids: a terminology muddle and a way out of it. Cell, 1987(50), 667. doi: 10.1016/0092-8674(87)90322-9
(2) Delsuc F, Brinkmann H, Philippe H. Phylogenomics and the reconstruction of the tree of life. Nat Rev Genet. 2005 May;6(5):361-75. Review. PubMed PMID: 15861208.
(3) Fitch W. (1970) Distinguishing homologous from analogous proteins. Systematic Zoology, 1970(19), 99. Retrieved from: http://www.jstor.org/stable/2412448
(4) Studer, R. A., Robinson-Rechavi, M. (2005)
How confident can we be that orthologs are similar, but paralogs differ? Trends in Genetics, 2005(25), 210. doi: 10.1016/j.tig.2009.03.004
“Homology” in proteins and nucleic acids: a terminology muddle and a way out of it. Cell, 1987(50), 667. doi: 10.1016/0092-8674(87)90322-9
(2) Delsuc F, Brinkmann H, Philippe H. Phylogenomics and the reconstruction of the tree of life. Nat Rev Genet. 2005 May;6(5):361-75. Review. PubMed PMID: 15861208.
(3) Fitch W. (1970) Distinguishing homologous from analogous proteins. Systematic Zoology, 1970(19), 99. Retrieved from: http://www.jstor.org/stable/2412448
(4) Studer, R. A., Robinson-Rechavi, M. (2005)
How confident can we be that orthologs are similar, but paralogs differ? Trends in Genetics, 2005(25), 210. doi: 10.1016/j.tig.2009.03.004